Clinical problem
Pneumothorax is a critical medical condition, commonly referred to as a collapsed lung. It can be painful and is potentially deadly, so pneumothorax requires immediate attention. Pneumothorax may be caused by an underlying lung disease, a blunt chest injury or can occur after surgery. It can also occur spontaneously. It is generally diagnosed by radiologists through examining chest radiographs (chest x-rays) and can be difficult to confirm - can you see the pneumothorax in the image shown here?! For further information see the Radiopaedia definition or the Radiology Assistant.
Recently, the Society for Imaging Informatics in Medicine (SIIM) organized a Kaggle challenge to tackle the detection and segmentation of this condition on chest x-ray images. We are interested in creating an open-source algorithm that uses this Kaggle dataset (initially) and other open datasets as well as chest x-ray data we have collected at Radboud UMC to detect pneumothorax.
Solution
Artificial Intelligence can be used to detect pneumothorax in chest x-rays automatically.
Research question and tasks
Detection of pneumothorax in chest x-rays.
Tasks
- Research previous work on segmentation of 2D medical images
- Research previous work on pneumothorax detection
- Develop a deep learning method that tackles the task
- Issue late submissions in the leaderboard and compare model performance with winning submissions
- Test the model on data from different institutions
- Publish the algorithm under an open-source license
Innovation
The goal is to integrate the algorithm in a set of chest x-ray analysis algorithms that are tested in routine clinical care at the Department of Radiology and Nuclear Medicine at Radboud UMC.
Results
The final thesis consists of three parts: 1. Open-source pneumothorax detection using deep learning 2. Unsupervised domain adaptation method: iterative self-training 3. Verify the approaches on our local private dataset: RadboudCXR
Open-source pneumothorax detection
In part I of this thesis, we show an open & transparent deep learning approach to pneumothorax detection. We combine various publicly available datasets that are accessible to anyone, giving us a verifiable multi-center approach. Even though we only train on public datasets, we achieve equal performance (AUC of 0.94) compared to state-of-the-art research that uses additional private datasets (Majkowska et al., 2020). Because all the datasets used in this research are public, we publish our model weights & algorithm and hope that future research can benefit from our work.
Our final models are trained on the public datasets SIIM, Chexpert, and MIMIC-CXR. Related work tends to struggle with the number of images containing a chest tube. We try to get rid of these chest tubes by selecting only the first study of a patient.
Careful hyperparameter optimization has been done (lung masking, normalizations, LR scheduling, data augmentation, etc.). One surprising result is that lung masking does not tend to work well for pneumothorax detection. This can be attributed to the fact that global image features such as the mediastinal shift and deep sulcus sign are cropped out by the lung masking algorithm.
Compared to related work, we achieve equal performance as related work by evaluating on a public dataset ChestXray14. Below we show the results.
| Research | # Pneumothorax | # Total | Model AUC |
|---|---|---|---|
| ChestXray14 (Wang et al., 2017) | 1.060 | 22.424 | 0.7993 |
| ChestXray14 (Taylor; Mielke, and Mongan, 2018) | 5.302 | 112.120 | 0.75 |
| ChestXray14 (Guendel et al., 2018) | 1.060 | 22.424 | 0.846 |
| ChestXray14 (Rajpurkar et al., 2018) | 45 | 420 | 0.944 |
| ChestXray14 (Majkowska et al., 2020) | 195 | 1.962 | 0.94 |
| Our research (EfficientNet-B3, 1024x1024) | 195 | 1.962 | 0.939 |
| Our research (EfficientNet-B3 + iterative self-training, 1024x1024) | 195 | 1.962 | 0.944 |
| Our research (DenseNet-121, 1024x1024) | 195 | 1.962 | 0.942 |
Radiologist performance
| Model | Specificity (%) | Sensitivity (%) | PPV (%) |
|---|---|---|---|
| Majkowska et al., 2020 | 90.8 | 72.8 | 48.7 |
| Our research (Densenet-121, 1024x1024) | 91.69 | 79.4 | 45.1 |
| Radiologists | 92.8 | 79.2 | 54.8 |
From these results, we can conclude that radiologists are still better at assessing the pneumothorax compared to our & related work, but the gap is getting closer.
Unsupervised Domain Adaptation: iterative self-training
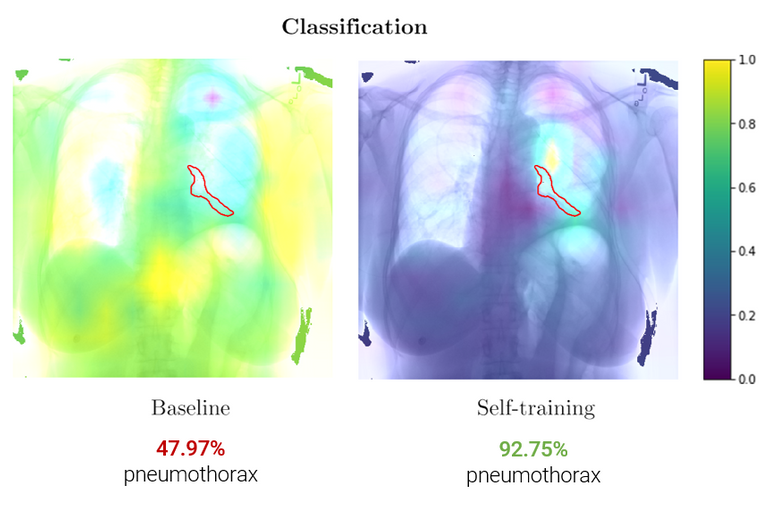
Further, in part II, we propose an unsupervised domain adaptation method – iterative self-training – that improves performance on an unseen dataset without the need for additional labelling (i.e. different hospital data). These results show an increase in performance (AUC 0.82 -> 0.89) for pneumothorax detection on public datasets CheXpert -> SIIM. This method was submitted to MIDL 2020 as a short paper.
These class activation maps show an increase in pneumothorax localization (red outline) after iterative self-training.
Clinical validation
Finally, in part III we evaluate the complete pipeline including our iterative self-training method on a local private dataset of 28.207 images (RadboudCXR) and verify that iterative self-training successfully adapts to an unseen local dataset (AUC 0.87 -> 0.92). We integrate the final algorithm using the grand-challenge platform and is publicly accessible for testing:
Example
Here we picked a random chest radiograph where our iterative self-training reaped benefits. For this case, we have a patient that just came from surgery and is currently in the intensive care unit. A confirmatory chest radiograph is requested to check whether the surgery caused any further complications.
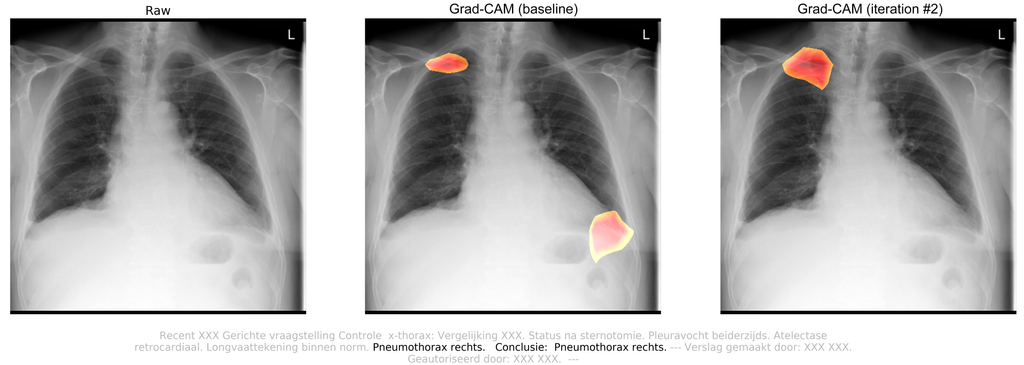
Attached radiology report:
Targeted questioning for a confirmatory chest radiograph. Status after sternotomy. Pleural effusions on both sides. Atelectasis retrocardiac. Lung vessel drawing within standards. Pneumothorax on the right side. Conclusion: Pneumothorax right lung.
By observing this chest radiograph, radiologists concluded that the patient suffered from a traumatic pneumothorax in the right lung, which was caused by surgery. After Iterative self-training, the algorithm successfully picked up the pneumothorax. The pneumothorax probability increased from 28.58% (FN) to 79.91% (TP). The class activation maps also show increased localization.