Background
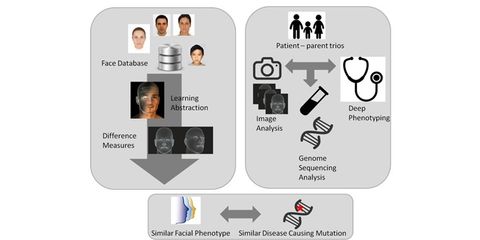
lntellectual disability (lD), affecting 2% of the general population, is frequently part of a syndrome with specific facial characteristics, so-called dysmorphism. The majority of lD cases are caused by rare sporadic genetic mutations, making it difficult to accurately diagnose. For clinical geneticists, craniofacial characteristics are highly informative when diagnosing genetic diseases. We propose to automate the extraction of patient-specific facial phenotypic information from image data. To this end, we will develop new machine learning algorithms for the analysis of facial expressions that are able to capture the relevant features for accurate matching of facial characteristics. This analysis will be supported by whole exome sequencing to assess the presence of a mutation in the patient. The main challenges in this study come from the novelty of the application area, the clinical conditions, and the rarity of some of the genetic disorders. Successful completion of the study will result in a technically and clinically validated technology that provides quick, automated, quantitative, and objective phenotyping of patients. The foreseen technology will reduce the burden on both the medical system and patient families by providing a quick non-invasive strategy able to increase the number of positive diagnoses of patients with lD.
Research question and tasks
Quantitative phenotyping of facial features can be regarded a challenging machine learning problem. lt requires (1) a meaningful characterization of facial features, (2) an ability to quantify the correspondences between multiple different faces, and (3) an expression of the probability that a patient's phenotypic appearance matches those associated to a known genetic disorder. We plan to address these problems using new advances in machine learning. Face representations will be estimated using deep neural networks (DNNs) that are trained on large datasets.
Tasks
- Development of deep neural networks for learning face representations
- Design of techniques to match faces to syndromes
- Analysis of learnt representations and resulting cluster assignments
Innovation
This project represents a unique combination of state-of-the-art genomic analysis, with significant developments in the phenotyping of patients with rare and difficult to diagnose developmental disorders via facial image analysis. This innovation comes through combining the two major research fields of genetics and machine learning, and requires a multidisciplinary team of clinicians, bioinformaticians and informaticians.
The developed tool for quantitative facial phenotyping will be tested and subsequently implemented in the Dept. of Human Genetics, Radboudumc. Further dissemination will follow both at national as well as international level.
Requirements
- Students in the final year of a Master study in computer science, artificial intelligence, biomedical engineering, or a related area.
Information
- Project duration: 6 months
- Location: Radboud University Medical Center
- For more information, please contact Luca Ambrogioni


